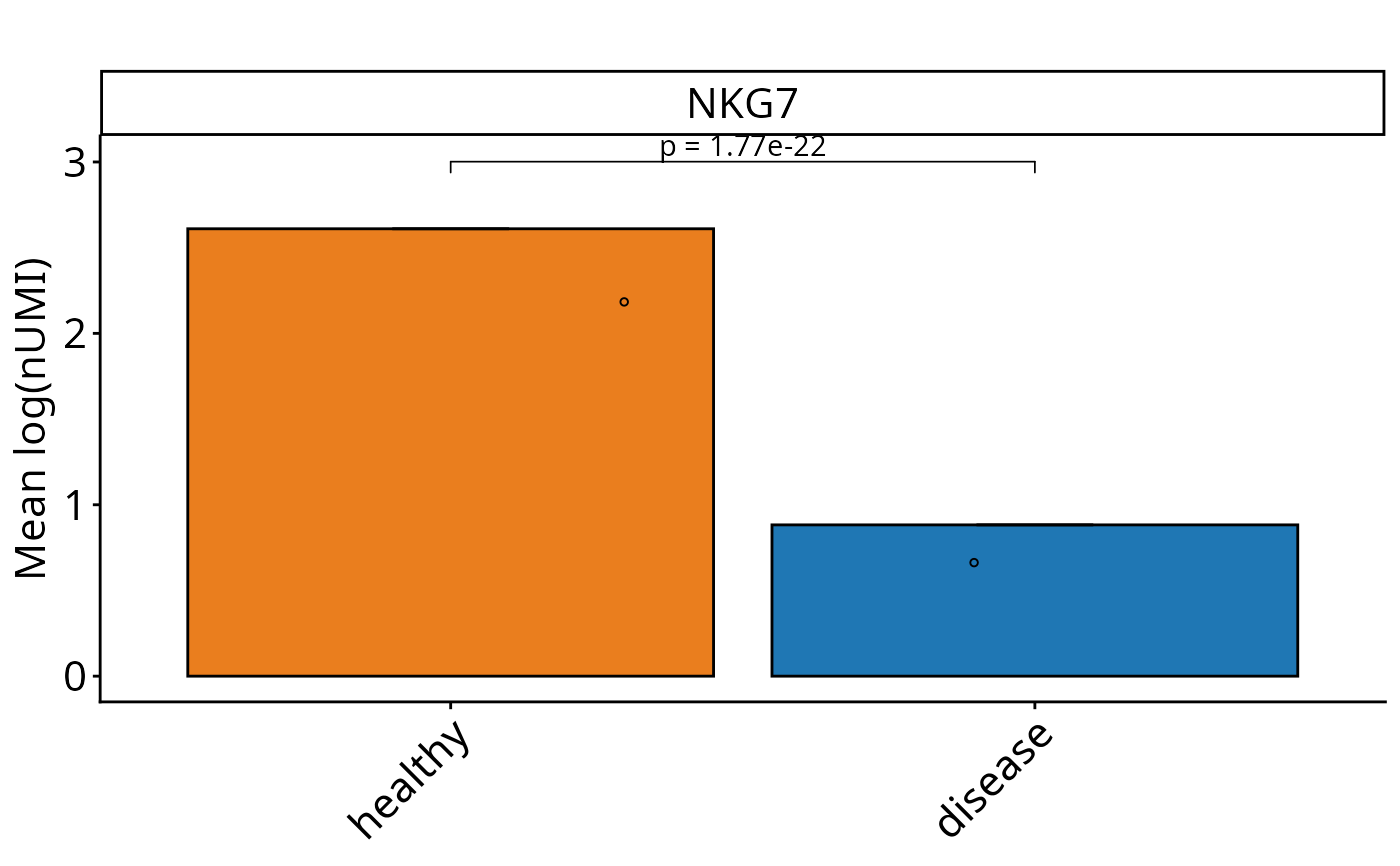
Perform SEM-based graphs with Wilcox test on single-cell level for Seurat and SCE objects. Calculates mean expression values and SEM for the selected feature, and visualizes them. Performs pairwise Wilcox tests comparing conditions, with optional custom control condition and clustering. Optionally returns a summary data frame, statistical test results, and the generated plot.
Usage
DO.Barplot(
sce_object,
Feature,
ListTest = NULL,
returnValues = FALSE,
ctrl.condition = NULL,
group.by = "condition",
test_use = "wilcox",
correction_method = "fdr",
p_values = NULL,
bar_colours = NULL,
stat_pos_mod = 1.15,
step_mod = 0.2,
x_label_rotation = 45,
plot_raw_pvalue = FALSE,
y_limits = NULL,
log1p_nUMI = TRUE,
random_seed = 42
)Arguments
- sce_object
combined SCE object or Seurat
- Feature
name of the feature/gene
- ListTest
List for which conditions wilcoxon test will be performed, if NULL always CTRL group against everything
- returnValues
return data frames needed for the plot, containing df.melt, df.melt.sum, df.melt.orig and wilcoxstats
- ctrl.condition
set your ctrl condition, relevant if running with empty comparison List
- group.by
select the seurat object slot where your conditions can be found, default conditon
- test_use
perform one of c( "wilcox", "wilcox_limma", "bimod", "t", "negbinom", "poisson", "LR", "MAST", "DESeq2", "none" ). default "wilcox"
- correction_method
correction for p-value calculation. One of c("BH", "bonferroni", "holm", "BY", "fdr", "none")
- p_values
Manually providing p-values for plotting, be aware of group size and if necessary make your test return the same amount of values
- bar_colours
colour vector
- stat_pos_mod
Defines the distance to the graphs of the statistic
- step_mod
Defines the distance between each statistics bracket
- x_label_rotation
Rotation of x-labels
- plot_raw_pvalue
plot the non adjusted p-value without correcting for multiple tests
- y_limits
set limits for y-axis
- log1p_nUMI
If nUMIs should be log1p transformed
- random_seed
parameter for random state initialisation
Examples
sce_data <-
readRDS(system.file("extdata", "sce_data.rds", package = "DOtools"))
ListTest <- list()
ListTest[[1]] <- c("healthy", "disease")
DO.Barplot(
sce_object = sce_data,
Feature = "NKG7",
test_use = "wilcox",
correction_method="fdr",
ListTest = ListTest,
ctrl.condition = "healthy",
group.by = "condition"
)
#> Using condition, orig.ident as id variables
#> For a (much!) faster implementation of the Wilcoxon Rank Sum Test,
#> (default method for FindMarkers) please install the presto package
#> --------------------------------------------
#> install.packages('devtools')
#> devtools::install_github('immunogenomics/presto')
#> --------------------------------------------
#> After installation of presto, Seurat will automatically use the more
#> efficient implementation (no further action necessary).
#> This message will be shown once per session